728x90
반응형

반응형
🙆♂️기본적인 코드 구조
from neuron import h, gui
from neuron.units import ms, mV
h.load_file('stdrun.hoc')
class Cell:
def __init__(self, gid, x, y, z, theta):
self._gid = gid
self._setup_morphology()
self.all = self.soma.wholetree()
self._setup_biophysics()
self.x = self.y = self.z = 0 # <-- NEW
h.define_shape()
self._rotate_z(theta) # <-- NEW
self._set_position(x, y, z) # <-- NEW
def __repr__(self):
return '{}[{}]'.format(self.name, self._gid)
# everything below here is NEW
def _set_position(self, x, y, z):
for sec in self.all:
for i in range(sec.n3d()):
sec.pt3dchange(i,
x - self.x + sec.x3d(i),
y - self.y + sec.y3d(i),
z - self.z + sec.z3d(i),
sec.diam3d(i))
self.x, self.y, self.z = x, y, z
def _rotate_z(self, theta):
"""Rotate the cell about the Z axis."""
for sec in self.all:
for i in range(sec.n3d()):
x = sec.x3d(i)
y = sec.y3d(i)
c = h.cos(theta)
s = h.sin(theta)
xprime = x * c - y * s
yprime = x * s + y * c
sec.pt3dchange(i, xprime, yprime, sec.z3d(i), sec.diam3d(i))
class BallAndStick(Cell):
name = 'BallAndStick'
def _setup_morphology(self):
self.soma = h.Section(name='soma', cell=self)
self.dend = h.Section(name='dend', cell=self)
self.dend.connect(self.soma)
self.soma.L = self.soma.diam = 12.6157
self.dend.L = 200
self.dend.diam = 1
def _setup_biophysics(self):
for sec in self.all:
sec.Ra = 100 # Axial resistance in Ohm * cm
sec.cm = 1 # Membrane capacitance in micro Farads / cm^2
self.soma.insert('hh')
for seg in self.soma:
seg.hh.gnabar = 0.12 # Sodium conductance in S/cm2
seg.hh.gkbar = 0.036 # Potassium conductance in S/cm2
seg.hh.gl = 0.0003 # Leak conductance in S/cm2
seg.hh.el = -54.3 # Reversal potential in mV
# Insert passive current in the dendrite
self.dend.insert('pas')
for seg in self.dend:
seg.pas.g = 0.001 # Passive conductance in S/cm2
seg.pas.e = -65 # Leak reversal potential mV일단 기본적으로 Cell 클래스와 BallAndStick 클래스 모습이 이렇습니다.
만드는 방법은 다른 함수를 통해서 만들어진 값들을 배열에 저장합니다.
def create_n_BallAndStick(n, r):
"""n = number of cells; r = radius of circle"""
cells = []
for i in range(n):
theta = i * 2 * h.PI / n
cells.append(BallAndStick(i, h.cos(theta) * r, h.sin(theta) * r, 0, theta))
return cellsmy_cells = create_n_BallAndStick(5, 50)위 함수를 사용해서 5개의 셀이 들어있는 배열을 만들 수 있고
stim = h.NetStim() # Make a new stimulator
# Attach it to a synapse in the middle of the dendrite
# of the first cell in the network. (Named 'syn_' to avoid
# being overwritten with the 'syn' var assigned later.)
syn_ = h.ExpSyn(my_cells[0].dend(0.5))
stim.number = 1
stim.start = 9
ncstim = h.NetCon(stim, syn_)
ncstim.delay = 1
ncstim.weight[0] = 0.04 # NetCon weight is a vector.
syn_.tau = 2NetStim을 활용하여 전기적 신호를 주입할 수 있습니다.
0번 셀의 dend 0.5부분을 ExpSyn 객체로 연결시켜서 syn_변수로 반환하였고
tau값을 설정했습니다. tau 값은 전류 감쇠에 영향을 줍니다.
recording_cell = my_cells[0]
soma_v = h.Vector().record(recording_cell.soma(0.5)._ref_v)
dend_v = h.Vector().record(recording_cell.dend(0.5)._ref_v)
t = h.Vector().record(h._ref_t)
h.finitialize(-65)
h.continuerun(25)%matplotlib inline
import matplotlib.pyplot as plt
plt.plot(t, soma_v, label='soma(0.5)')
plt.plot(t, dend_v, label='dend(0.5)')
plt.legend()
plt.show()기록하고 plot하는 코드입니다. 이제 다양한 부분들을 확인해보겠습니다.
🙋♂️ 여러 dendrite 붙이기

기존 코드의 모습은 이렇습니다.
class Cell:
def __init__(self, gid, x, y, z, theta):
self._gid = gid
self._setup_morphology()
self.all = self.soma.wholetree()
self._setup_biophysics()
self.x = self.y = self.z = 0 # <-- NEW
h.define_shape()
self._rotate_z(theta) # <-- NEW
self._set_position(x, y, z) # <-- NEW
def __repr__(self):
return '{}[{}]'.format(self.name, self._gid)
# everything below here is NEW
def _set_position(self, x, y, z):
for sec in self.all:
for i in range(sec.n3d()):
sec.pt3dchange(i,
x - self.x + sec.x3d(i),
y - self.y + sec.y3d(i),
z - self.z + sec.z3d(i),
sec.diam3d(i))
self.x, self.y, self.z = x, y, z
def _rotate_z(self, theta):
"""Rotate the cell about the Z axis."""
for sec in self.all:
for i in range(sec.n3d()):
x = sec.x3d(i)
y = sec.y3d(i)
c = h.cos(theta)
s = h.sin(theta)
xprime = x * c - y * s
yprime = x * s + y * c
sec.pt3dchange(i, xprime, yprime, sec.z3d(i), sec.diam3d(i))
class BallAndStick(Cell):
name = 'BallAndStick'
def _setup_morphology(self):
self.soma = h.Section(name='soma', cell=self)
self.dend1 = h.Section(name='dend1', cell=self)
self.dend2 = h.Section(name='dend2', cell=self)
self.dend1.connect(self.soma)
self.dend2.connect(self.soma)
self.soma.L = self.soma.diam =12.6157
self.dend1.L = self.dend2.L= 200
self.dend1.diam = self.dend2.diam = 1
def _setup_biophysics(self):
for sec in self.all:
sec.Ra = 100 # Axial resistance in Ohm * cm
sec.cm = 1 # Membrane capacitance in micro Farads / cm^2
self.soma.insert('hh')
for seg in self.soma:
seg.hh.gnabar = 0.12 # Sodium conductance in S/cm2
seg.hh.gkbar = 0.036 # Potassium conductance in S/cm2
seg.hh.gl = 0.0003 # Leak conductance in S/cm2
seg.hh.el = -54.3 # Reversal potential in mV
# Insert passive current in the dendrite
self.dend1.insert('pas')
for seg in self.dend1:
seg.pas.g = 0.001 # Passive conductance in S/cm2
seg.pas.e = -65 # Leak reversal potential mV
self.dend2.insert('pas')
for seg in self.dend2:
seg.pas.g = 0.001 # Passive conductance in S/cm2
seg.pas.e = -65 # Leak reversal potential mVdendrite 2개를 붙여서 수정한 모습은 이렇습니다! 잘 연결된 모습입니다.
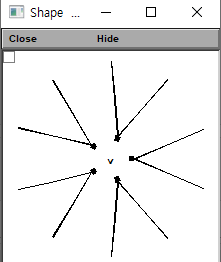
시뮬레이션 돌리고 plot 해보기
recording_cell = my_cells[0]
soma_v = h.Vector().record(recording_cell.soma(0.5)._ref_v)
dend_v1 = h.Vector().record(recording_cell.dend1(0.5)._ref_v)
dend_v2 = h.Vector().record(recording_cell.dend2(0.5)._ref_v)
t = h.Vector().record(h._ref_t)
h.finitialize(-65)
h.continuerun(25)
%matplotlib inline
import matplotlib.pyplot as plt
plt.plot(t, soma_v, label='soma(0.5)')
plt.plot(t, dend_v1, label='dend(0.5)')
plt.plot(t, dend_v2, label='dend(0.5)')
plt.legend()
plt.show()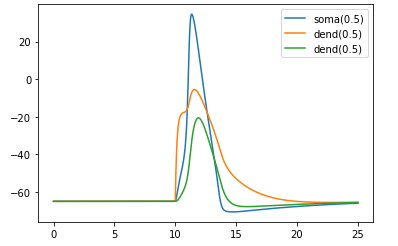
흥미로운 결과값이 나왔습니다.
728x90
반응형
'AI > neuron' 카테고리의 다른 글
| [neuron][파이썬] 19. 뉴런 병렬 통신 - Parallel communication in NEURON (0) | 2022.07.28 |
|---|---|
| [neuron][파이썬] 21. 깃헙 뉴런 튜토리얼 - Ball and Stick model (0) | 2022.07.27 |
| [neuron][파이썬] 21. 깃헙 뉴런 튜토리얼 - Single compartment neuron model (0) | 2022.07.27 |
| [neuron][파이썬] 18. 확장된 링 네트워크 (0) | 2022.07.25 |
| [neuron][파이썬] 17. 링 네트워크 시뮬레이션 실행 및 출력 (0) | 2022.07.22 |